SASA Validation
Accuracy comparison of zsasa against reference implementations across the E. coli K-12 proteome (4,370 structures).
Note: Validation uses batch processing across the entire proteome. MD trajectory validation is covered in md.md.
TL;DR

| Algorithm | Tool | R² | Mean Error % | Max Error % |
|---|---|---|---|---|
| SR (100 pts) | zsasa f64 | 1.000000 | 0.0000 | 0.0000 |
| SR (100 pts) | zsasa f32 | 1.000000 | 0.0001 | 0.0150 |
| SR (100 pts) | zsasa bitmask | 0.999721 | 0.81 | 2.51 |
| SR (100 pts) | RustSASA | 0.999963 | 0.32 | 2.49 |
| SR (128 pts) | Lahuta bitmask | 0.999768 | 0.73 | 2.47 |
| LR (20 slices) | zsasa f64 | 0.999980 | 0.22 | 0.31 |
- zsasa f64 is bit-identical to FreeSASA at all point counts
- zsasa f32 has negligible rounding error (max 0.015%)
- Bitmask error plateaus at ~0.7–0.8% (LUT approximation, independent of point count)
- RustSASA converges with point count: 0.32% at 100 → 0.06% at 1000
- Lahuta bitmask (128 pts): comparable accuracy to zsasa bitmask
Test Environment
| Item | Value |
|---|---|
| Machine | MacBook Pro |
| Chip | Apple M4 (10 cores: 4P + 6E) |
| Memory | 32 GB |
| OS | macOS 15.3.2 (Darwin 24.6.0) |
Shrake-Rupley (SR)
Dataset: AlphaFold E. coli K-12 proteome, 4,370 structures. Reference: FreeSASA.
n_points Convergence
| Tool | n_points | N | R² | Mean Error % | Max Error % |
|---|---|---|---|---|---|
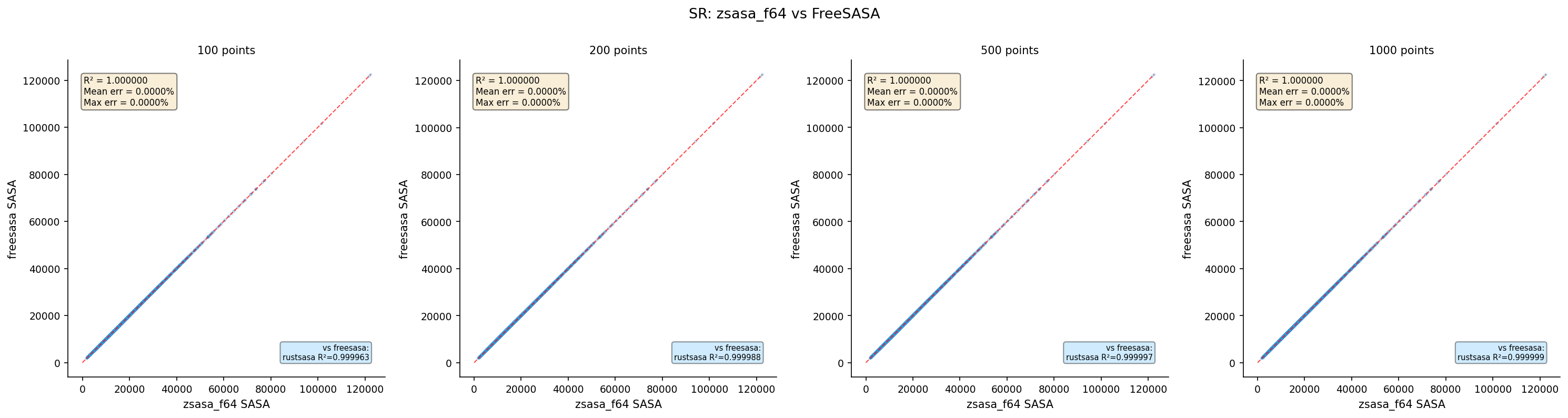
| zsasa f64 | 100 | 4,370 | 1.000000 | 0.0000 | 0.0000 |
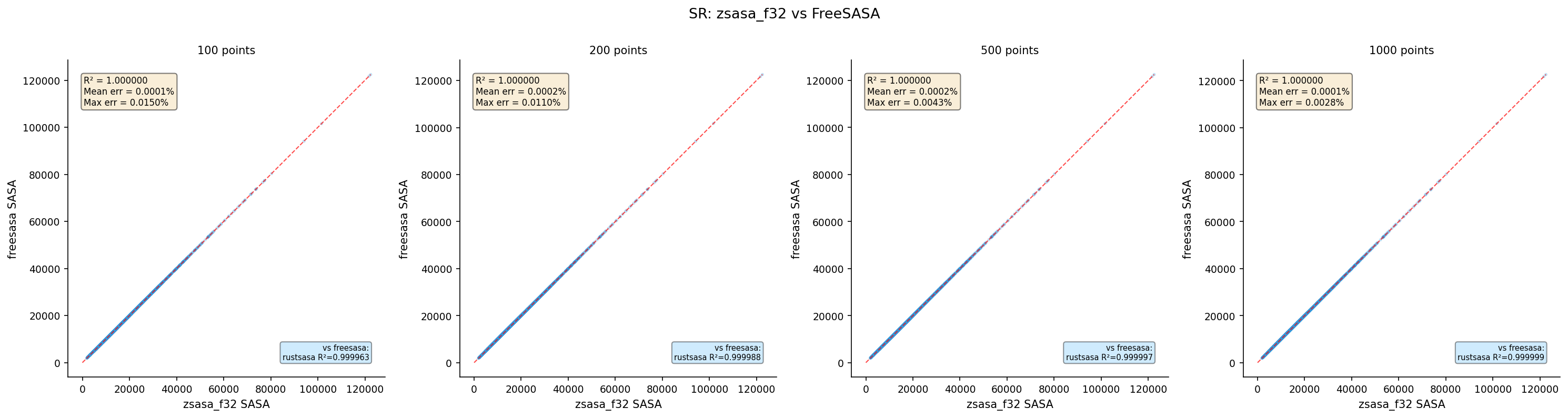
| zsasa f32 | 100 | 4,370 | 1.000000 | 0.0001 | 0.0150 |
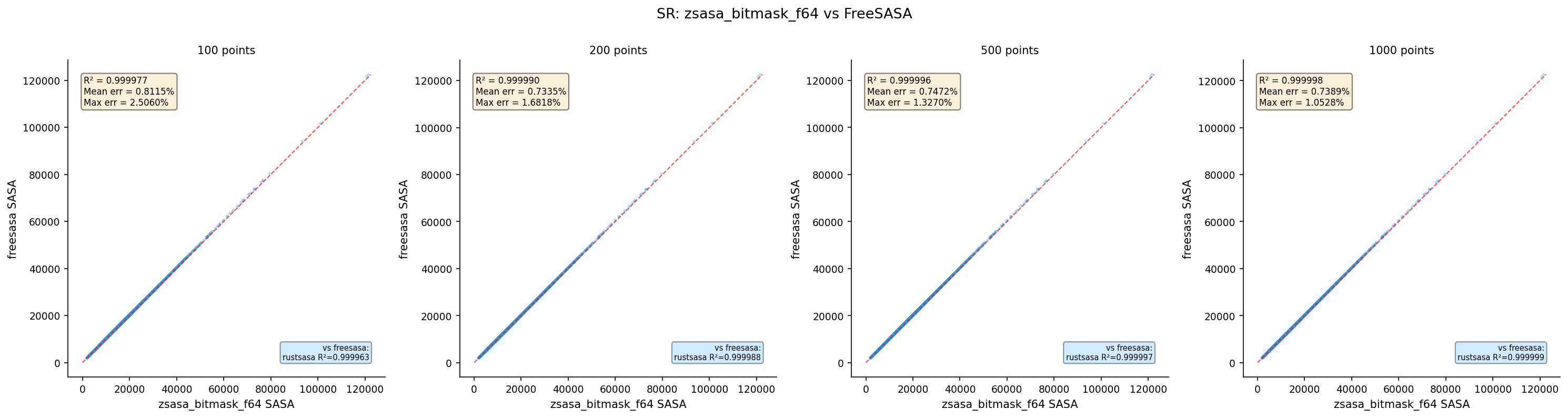
| zsasa bitmask f64 | 100 | 4,370 | 0.999721 | 0.8115 | 2.5060 |
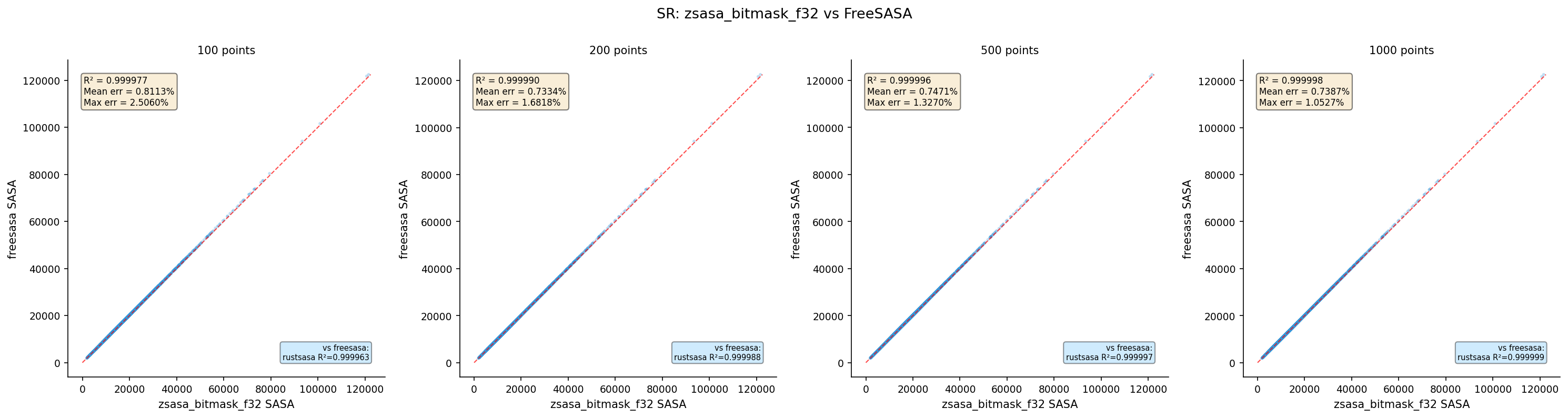
| zsasa bitmask f32 | 100 | 4,370 | 0.999721 | 0.8113 | 2.5060 |
| RustSASA | 100 | 4,370 | 0.999963 | 0.3181 | 2.4859 |
| zsasa f64 | 200 | 4,370 | 1.000000 | 0.0000 | 0.0000 |
| zsasa f32 | 200 | 4,370 | 1.000000 | 0.0002 | 0.0110 |
| zsasa bitmask f64 | 200 | 4,370 | 0.999781 | 0.7335 | 1.6818 |
| zsasa bitmask f32 | 200 | 4,370 | 0.999781 | 0.7334 | 1.6818 |
| RustSASA | 200 | 4,370 | 0.999988 | 0.1841 | 1.2255 |
| zsasa f64 | 500 | 4,370 | 1.000000 | 0.0000 | 0.0000 |
| zsasa f32 | 500 | 4,370 | 1.000000 | 0.0002 | 0.0043 |
| zsasa bitmask f64 | 500 | 4,370 | 0.999778 | 0.7472 | 1.3270 |
| zsasa bitmask f32 | 500 | 4,370 | 0.999778 | 0.7471 | 1.3270 |
| RustSASA | 500 | 4,370 | 0.999997 | 0.0923 | 0.5562 |
| zsasa f64 | 1000 | 4,370 | 1.000000 | 0.0000 | 0.0000 |
| zsasa f32 | 1000 | 4,370 | 1.000000 | 0.0001 | 0.0028 |
| zsasa bitmask f64 | 1000 | 4,370 | 0.999784 | 0.7389 | 1.0528 |
| zsasa bitmask f32 | 1000 | 4,370 | 0.999785 | 0.7387 | 1.0527 |
| RustSASA | 1000 | 4,370 | 0.999999 | 0.0558 | 0.3401 |
Key findings:
- zsasa f64: Bit-identical to FreeSASA at all point counts (same algorithm parameters)
- zsasa f32: Max 0.015% error from floating-point rounding — negligible for practical use
- Bitmask variants: Mean error ~0.7–0.8%, plateaus regardless of point count (LUT approximation error)
- RustSASA: Converges toward FreeSASA with increasing points (0.32% → 0.06% mean error)
- f32 vs f64 bitmask: Virtually identical — bitmask error dominates floating-point error
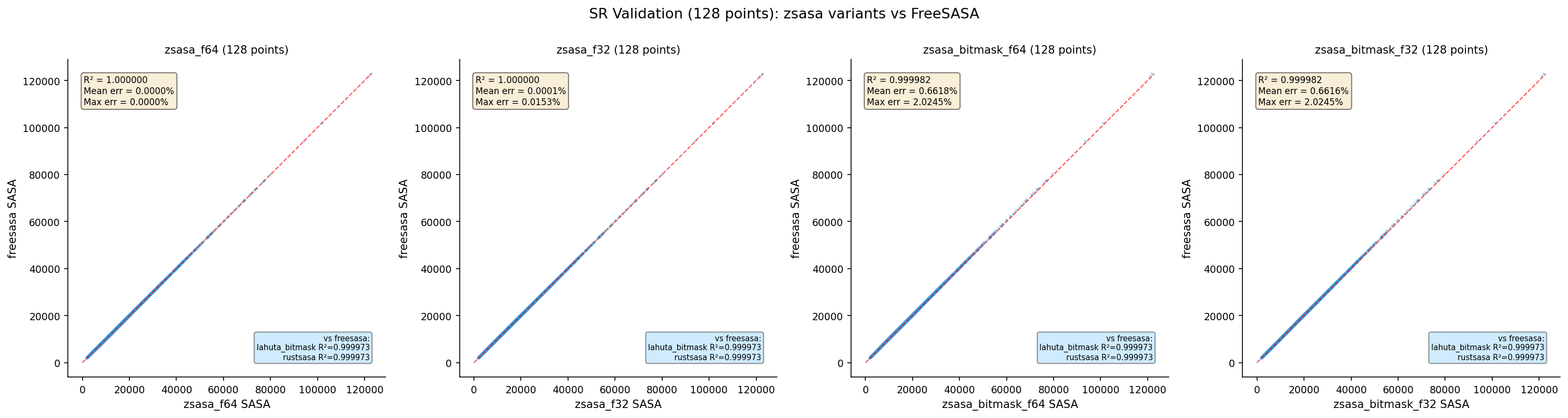
Lahuta Bitmask (128 pts)
Lahuta requires n_points=128 for bitmask support.
| Tool | N | R² | Mean Error % | Max Error % |
|---|---|---|---|---|
| zsasa f64 | 4,370 | 1.000000 | 0.0000 | 0.0000 |
| zsasa f32 | 4,370 | 1.000000 | 0.0001 | 0.0153 |
| zsasa bitmask f64 | 4,370 | 0.999811 | 0.6618 | 2.0245 |
| zsasa bitmask f32 | 4,370 | 0.999811 | 0.6616 | 2.0245 |
| RustSASA | 4,370 | 0.999973 | 0.2752 | 2.1487 |
| Lahuta bitmask | 4,370 | 0.999768 | 0.7316 | 2.4709 |
- Lahuta bitmask accuracy is comparable to zsasa bitmask (~0.73% vs ~0.66% mean error)
- Both use LUT bitmask neighbor lists — the error pattern is similar
Validation Plots

| zsasa f64 | zsasa f32 |
|---|---|
 |  |
| zsasa bitmask f64 | zsasa bitmask f32 |
|---|---|
 |  |
| Lahuta bitmask |
|---|
 |
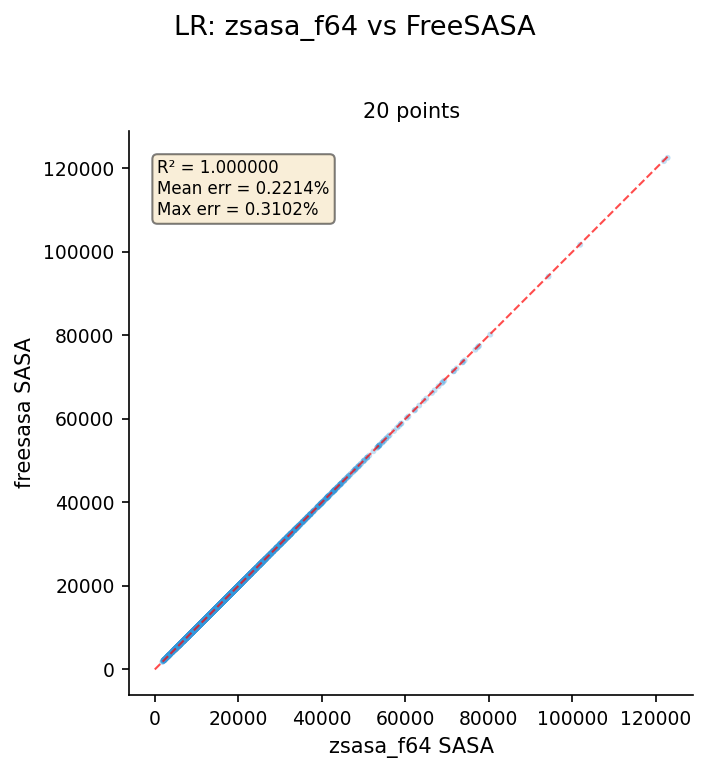
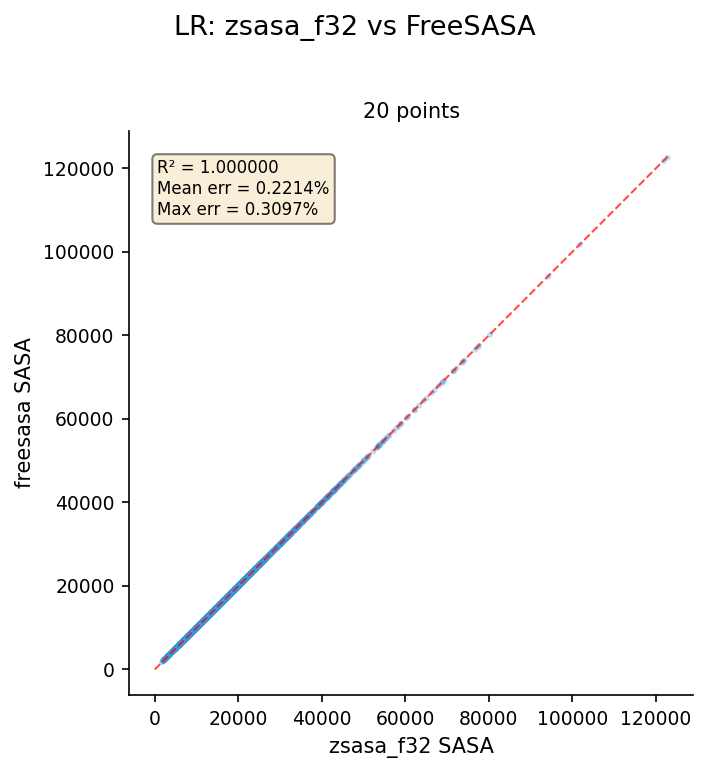
Lee-Richards (LR)
Dataset: AlphaFold E. coli K-12 proteome, 4,370 structures, n_slices=20. Reference: FreeSASA.
| Tool | N | R² | Mean Error % | Max Error % |
|---|---|---|---|---|
| zsasa f64 | 4,370 | 0.999980 | 0.2214 | 0.3102 |
| zsasa f32 | 4,370 | 0.999980 | 0.2214 | 0.3097 |
Key findings:
- f32 and f64 are identical — LR error comes from slice discretization differences, not floating-point precision
- Mean error ~0.22% is higher than SR (<0.001%) due to different slicing implementations between zsasa and FreeSASA
- R² = 0.999980 confirms strong linear agreement
Validation Plots

| zsasa f64 | zsasa f32 |
|---|---|
 |  |
Running Validation
# Shrake-Rupley (all tools, multi-point convergence)
./benchmarks/scripts/validation.py run \
-i benchmarks/UP000000625_83333_ECOLI_v6/pdb \
-n ecoli --algorithm sr
# -> benchmarks/results/validation/ecoli/sr/
# Lee-Richards
./benchmarks/scripts/validation.py run \
-i benchmarks/UP000000625_83333_ECOLI_v6/pdb \
-n ecoli --algorithm lr
# -> benchmarks/results/validation/ecoli/lr/
# Re-analyze existing results
./benchmarks/scripts/validation.py compare \
-d benchmarks/results/validation/ecoli/sr
Related Documents
- Single-File Benchmarks — Per-structure performance across 2,013 structures
- Batch Processing Benchmarks — Proteome-scale throughput
- MD Trajectory Benchmarks — MD trajectory performance and validation