Batch Processing Benchmarks
Throughput benchmarks for processing complete proteome datasets using hyperfine timing.
Note: Measures total wall-clock time for batch processing a directory of PDB files. All tools use native multi-threading. Test points: 128 (required for Lahuta bitmask support).
TL;DR
| Dataset | Structures | zsasa_bitmask (f32) | vs FreeSASA | vs RustSASA | vs Lahuta BM | RSS |
|---|---|---|---|---|---|---|
| E. coli | 4,370 | 1.42s | 8.9x faster | 3.7x faster | 1.4x faster | 43 MB |
| Human | 23,586 | 14.04s | 9.9x faster | 3.9x faster | 1.6x faster | 73 MB |
| SwissProt | 550,122 | 4m 02s | 8.0x faster | 2.7x faster | 1.3x faster | 157 MB |
- zsasa_bitmask (LUT bitmask) is the fastest tool across all datasets, using 3.7–7.2x less memory than RustSASA
- Memory stays flat (~157 MB) even at 550K structures, while RustSASA scales to 1.1 GB and Lahuta BM to 2.2 GB
Test Environment
| Item | Value |
|---|---|
| Machine | MacBook Pro |
| Chip | Apple M4 (10 cores: 4P + 6E) |
| Memory | 32 GB |
| OS | macOS 15.3.2 (Darwin 24.6.0) |
SwissProt additionally tested on M2 Max (96 GB) — see SwissProt section.
Tools Compared
| Tool | Language | Description |
|---|---|---|
| zsasa | Zig | f64/f32 precision, standard neighbor lists |
| zsasa_bitmask | Zig | f64/f32 precision, LUT bitmask neighbor lists |
| Lahuta | C++ | Standard neighbor lists |
| Lahuta Bitmask | C++ | LUT bitmask neighbor lists |
| RustSASA | Rust | Native multi-threading |
| FreeSASA | C | No native batch mode — custom wrapper (freesasa_batch.cc) |
E. coli Proteome (4,370 structures)
Dataset: AlphaFold E. coli K-12 proteome (UP000000625_83333_ECOLI_v6), PDB format.
10-Thread Comparison
Benchmark: warmup=3, runs=10, threads=10.
| Tool | Time (s) | Std Dev | files/sec | vs FreeSASA | vs RustSASA | vs Lahuta BM | RSS (MB) |
|---|---|---|---|---|---|---|---|
| zsasa_bitmask (f32) | 1.42 | ±0.013 | 3,085 | 8.9x | 3.7x | 1.4x | 43 |
| zsasa_bitmask (f64) | 1.42 | ±0.008 | 3,072 | 8.9x | 3.7x | 1.4x | 45 |
| Lahuta Bitmask | 2.01 | ±0.004 | 2,172 | 6.3x | 2.6x | baseline | 291 |
| zsasa (f32) | 4.09 | ±0.346 | 1,069 | 3.1x | 1.3x | 0.5x | 40 |
| zsasa (f64) | 4.08 | ±0.034 | 1,071 | 3.1x | 1.3x | 0.5x | 43 |
| RustSASA | 5.24 | ±0.035 | 834 | 2.4x | baseline | 0.4x | 169 |
| Lahuta | 6.70 | ±0.032 | 652 | 1.9x | 0.8x | 0.3x | 315 |
| FreeSASA | 12.60 | ±0.049 | 347 | baseline | 0.4x | 0.2x | 467 |
Key findings:
- zsasa_bitmask (f32) processes 4,370 structures in 1.42s — 3.7x faster than RustSASA, 1.4x faster than Lahuta Bitmask
- Memory: zsasa uses 40–45 MB vs RustSASA 169 MB (3.8x less), Lahuta BM 291 MB (6.5x less)
Thread Scaling
Benchmark: warmup=3, runs=3, threads=1,8,10.
| 1 thread | 8 threads | 10 threads |
|---|---|---|
 | 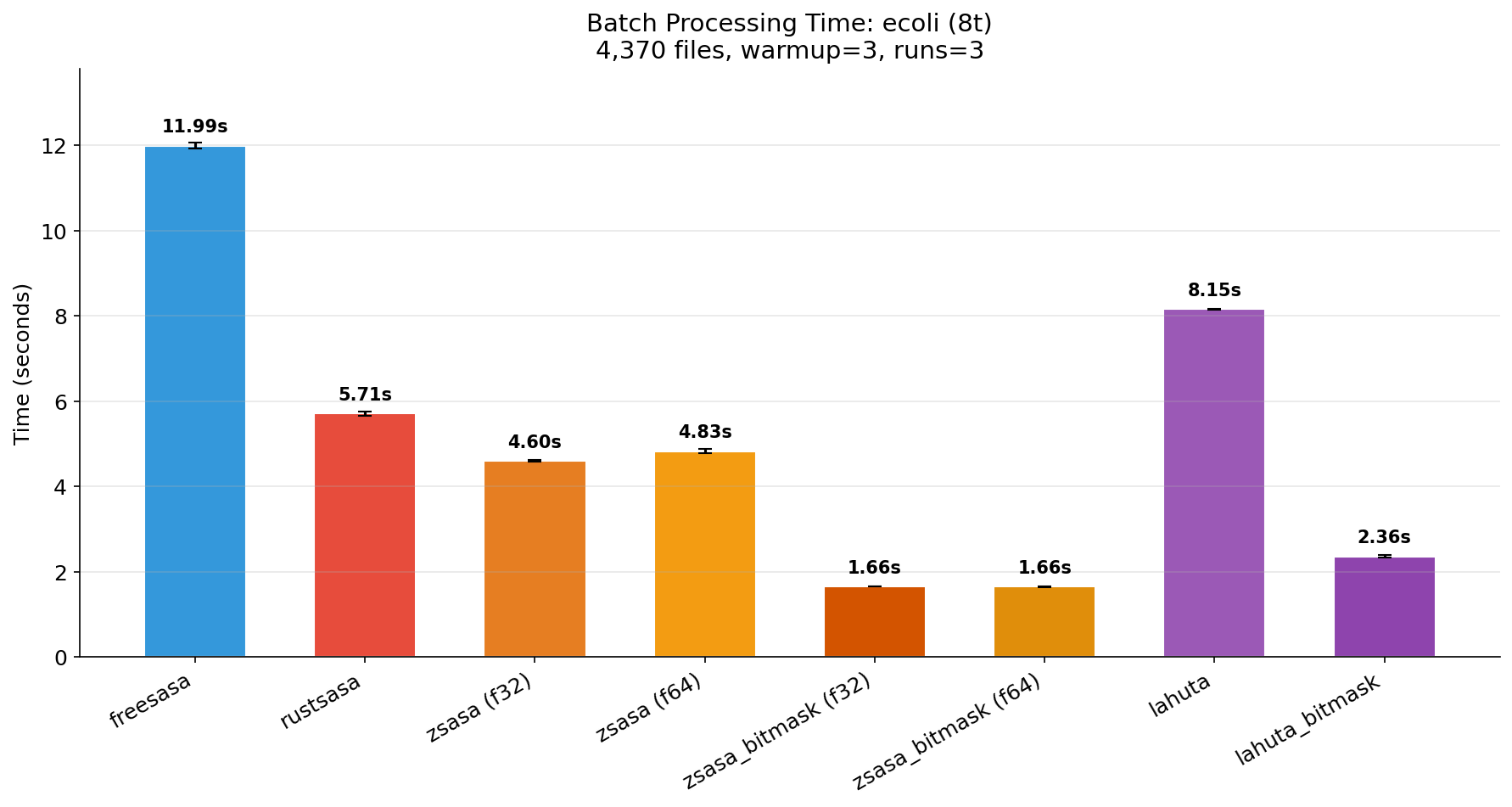 |  |
| Memory (1t) | Memory (8t) | Memory (10t) |
|---|---|---|
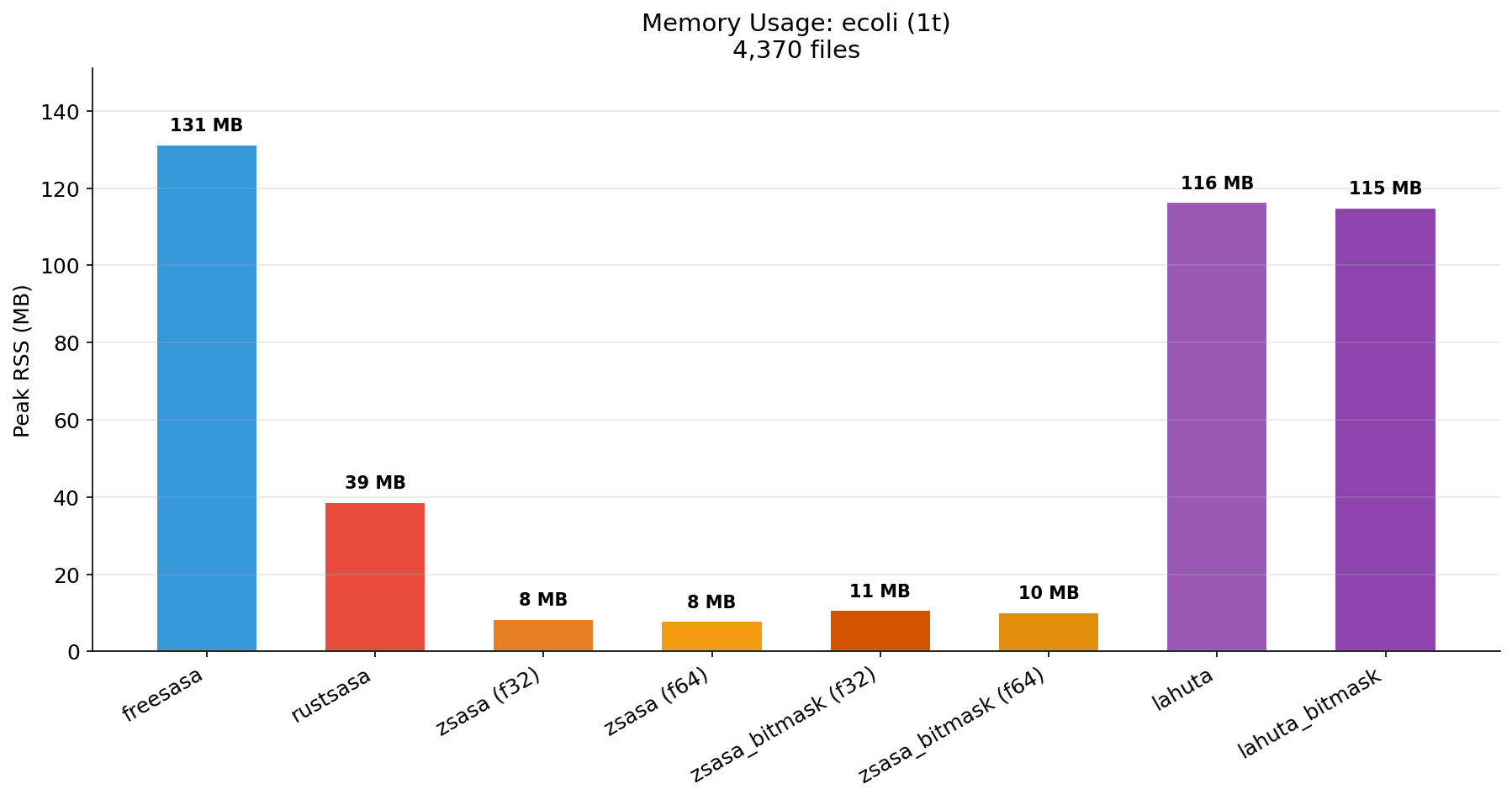 |  |  |
Human Proteome (23,586 structures)
Dataset: AlphaFold Human proteome (UP000005640_9606_HUMAN_v6), PDB format.
Benchmark: warmup=3, runs=10, threads=10.
| Tool | Time (s) | Std Dev | files/sec | vs FreeSASA | vs RustSASA | vs Lahuta BM | RSS (MB) |
|---|---|---|---|---|---|---|---|
| zsasa_bitmask (f32) | 14.04 | ±0.073 | 1,680 | 9.9x | 3.9x | 1.6x | 73 |
| zsasa_bitmask (f64) | 14.85 | ±0.579 | 1,589 | 9.3x | 3.6x | 1.5x | 77 |
| Lahuta Bitmask | 22.67 | ±0.250 | 1,040 | 6.1x | 2.4x | baseline | 1,415 |
| zsasa (f32) | 39.82 | ±0.997 | 592 | 3.5x | 1.4x | 0.6x | 70 |
| zsasa (f64) | 42.33 | ±1.483 | 557 | 3.3x | 1.3x | 0.5x | 75 |
| RustSASA | 54.16 | ±0.307 | 435 | 2.6x | baseline | 0.4x | 334 |
| Lahuta | 78.64 | ±0.674 | 300 | 1.8x | 0.7x | 0.3x | 1,077 |
| FreeSASA | 138.77 | ±0.664 | 170 | baseline | 0.4x | 0.2x | 1,913 |
Key findings:
- zsasa_bitmask (f32) processes 23,586 structures in 14.04s — 3.9x faster than RustSASA, 1.6x faster than Lahuta Bitmask
- Memory: zsasa uses 70–77 MB vs RustSASA 334 MB (4.5x less), Lahuta BM 1,415 MB (19x less)
| Time (10t) | Memory (10t) |
|---|---|
 | 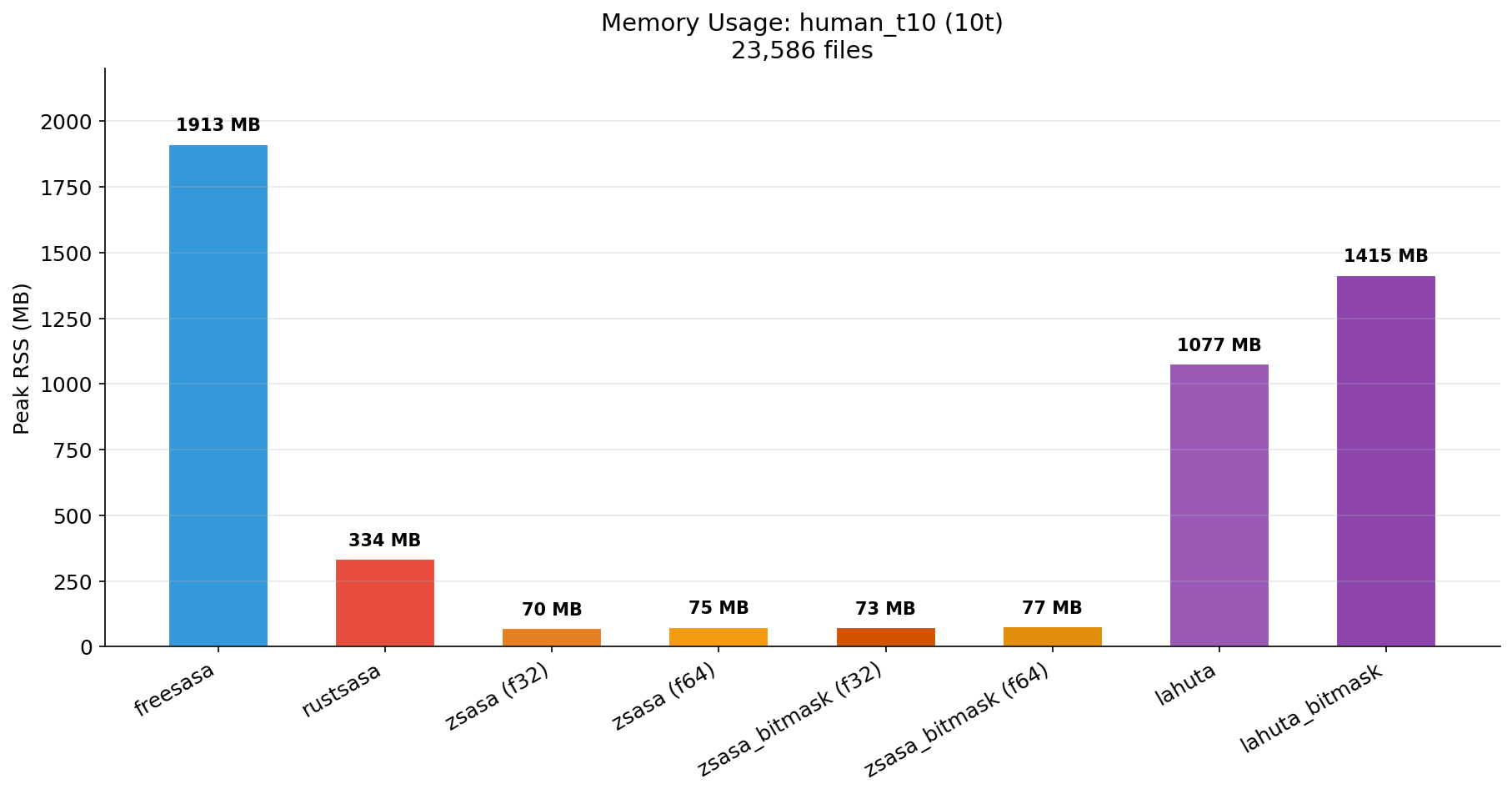 |
SwissProt (550,122 structures)
Dataset: SwissProt PDB v6, PDB format. The largest benchmark at 550K structures.
M2 Max (96 GB) — Compute Bound
| Item | Value |
|---|---|
| Chip | Apple M2 Max (12 cores) |
| Memory | 96 GB |
With sufficient RAM, all file data stays in the OS page cache and performance is purely compute-bound.
Benchmark: warmup=3, runs=3, threads=10.
| Tool | Time | Std Dev | files/sec | vs FreeSASA | vs RustSASA | vs Lahuta BM | RSS (MB) |
|---|---|---|---|---|---|---|---|
| zsasa_bitmask (f32) | 4m 02s | ±1.86 | 2,269 | 8.0x | 2.7x | 1.3x | 157 |
| zsasa_bitmask (f64) | 4m 07s | ±1.49 | 2,229 | 7.9x | 2.7x | 1.3x | 162 |
| Lahuta Bitmask | 5m 12s | ±3.32 | 1,761 | 6.2x | 2.1x | baseline | 2,187 |
| zsasa (f32) | 10m 39s | ±0.57 | 861 | 3.0x | 1.03x | 0.5x | 154 |
| zsasa (f64) | 10m 41s | ±1.43 | 858 | 3.0x | 1.03x | 0.5x | 159 |
| RustSASA | 10m 58s | ±1.30 | 835 | 2.9x | baseline | 0.5x | 1,131 |
| Lahuta | 16m 34s | ±2.51 | 553 | 2.0x | 0.66x | 0.3x | 1,873 |
| FreeSASA | 32m 21s | ±8.95 | 283 | baseline | 0.34x | 0.2x | 2,875 |
| Time (10t) | Memory (10t) |
|---|---|
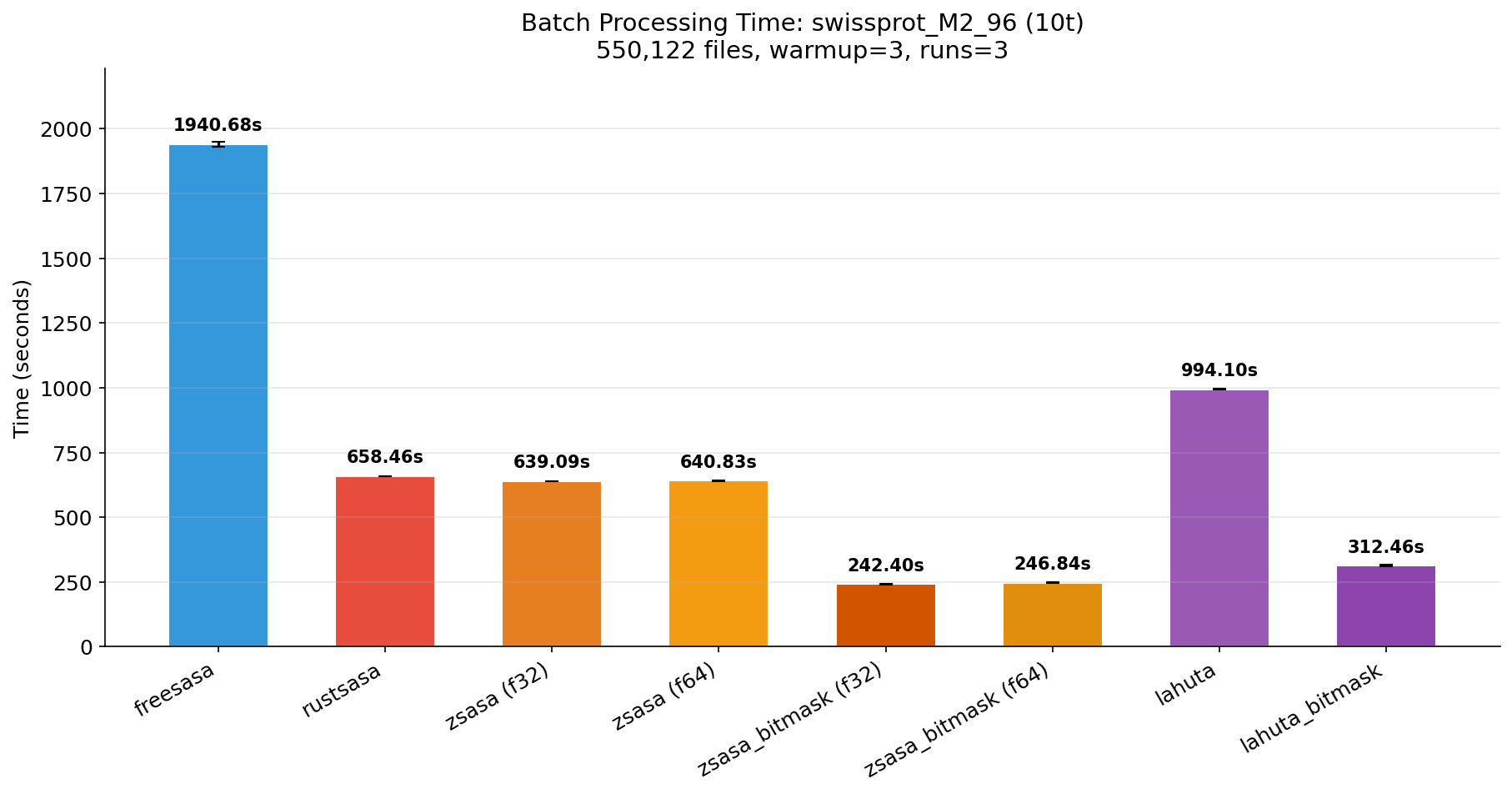 |  |
M4 (32 GB) — I/O Bound (mmap)
When the dataset exceeds available RAM, mmap page faults become the bottleneck and performance converges across tools.
Benchmark: warmup=3, runs=3, threads=10.
| Tool | Time | Std Dev | files/sec | vs FreeSASA | vs RustSASA | vs Lahuta BM | RSS (MB) |
|---|---|---|---|---|---|---|---|
| zsasa_bitmask (f32) | 11m 05s | ±6.48 | 828 | 2.9x | 2.4x | 1.0x | 157 |
| zsasa_bitmask (f64) | 11m 07s | ±2.14 | 824 | 2.9x | 2.4x | 1.0x | 161 |
| Lahuta Bitmask | 11m 08s | ±9.91 | 823 | 2.8x | 2.4x | baseline | 2,152 |
| zsasa (f32) | 16m 02s | ±11.87 | 572 | 2.0x | 1.6x | 0.7x | 154 |
| zsasa (f64) | 16m 11s | ±3.65 | 567 | 2.0x | 1.6x | 0.7x | 159 |
| Lahuta | 22m 11s | ±5.21 | 413 | 1.4x | 1.2x | 0.5x | 1,820 |
| RustSASA | 26m 16s | ±8.05 | 349 | 1.2x | baseline | 0.4x | 1,131 |
| FreeSASA | 31m 42s | ±7.67 | 289 | baseline | 0.8x | 0.4x | 2,440 |
| Time (10t) | Memory (10t) |
|---|---|
 | 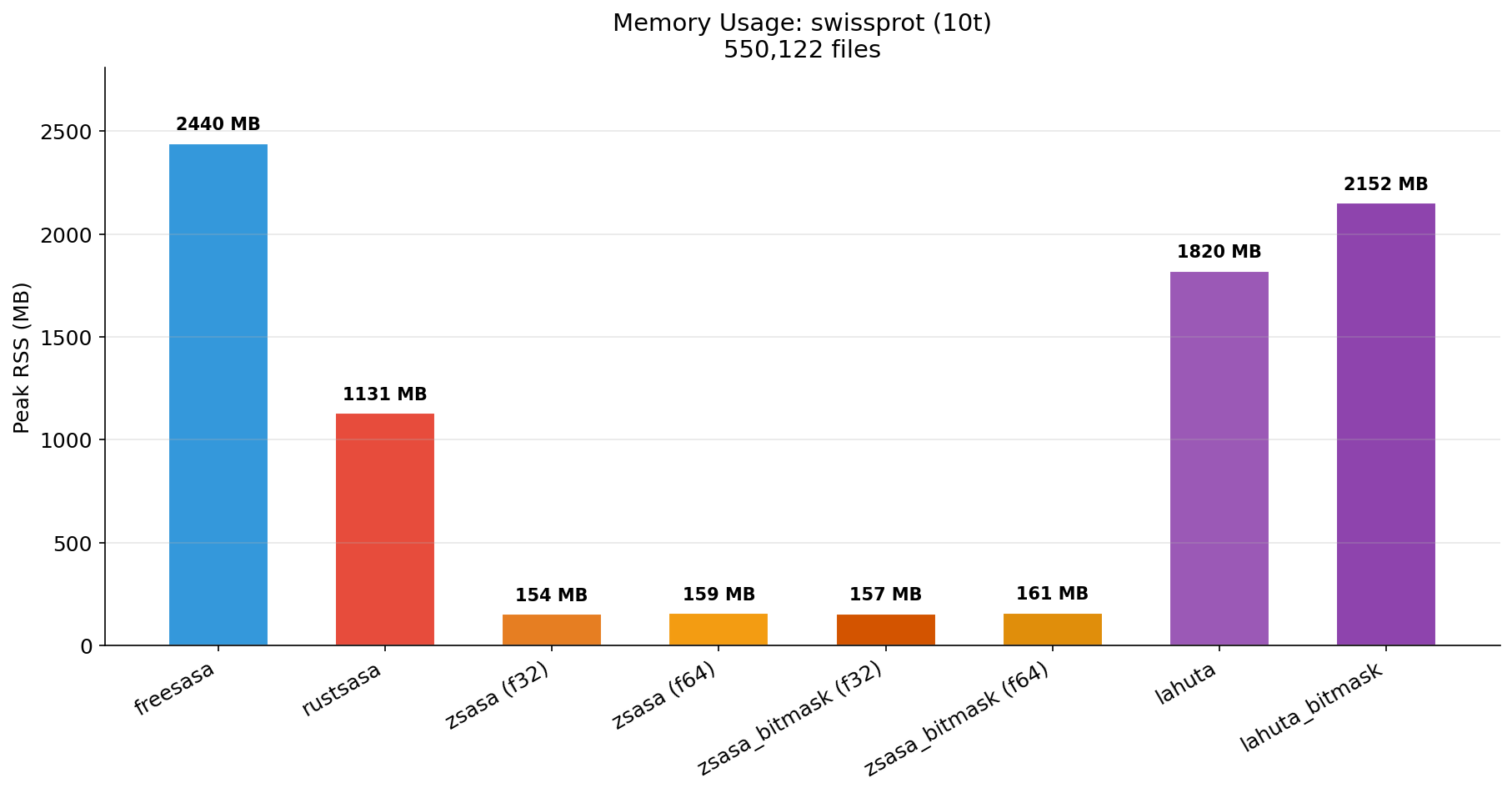 |
Key findings:
- M2 Max (96 GB): zsasa_bitmask completes 550K structures in 4 minutes — 2.7x faster than RustSASA, 1.3x faster than Lahuta BM
- M4 (32 GB): I/O-bound — zsasa_bitmask and Lahuta Bitmask converge at ~11 min (both 2.4x faster than RustSASA)
- Memory: zsasa uses ~157 MB regardless of dataset size, vs RustSASA 1.1 GB (7.2x less), Lahuta BM 2.2 GB (14x less)
Summary
Performance (10 threads)
| Dataset | Structures | zsasa_bitmask (f32) | vs RustSASA | vs Lahuta BM | RSS |
|---|---|---|---|---|---|
| E. coli | 4,370 | 1.42s | 3.7x | 1.4x | 43 MB |
| Human | 23,586 | 14.04s | 3.9x | 1.6x | 73 MB |
| SwissProt (96 GB) | 550,122 | 4m 02s | 2.7x | 1.3x | 157 MB |
| SwissProt (32 GB) | 550,122 | 11m 05s | 2.4x | 1.0x | 157 MB |
Memory Efficiency (10 threads)
| Dataset | zsasa | RustSASA | Lahuta BM | zsasa ratio |
|---|---|---|---|---|
| E. coli | 43 MB | 169 MB | 291 MB | 3.9x less than RustSASA |
| Human | 73 MB | 334 MB | 1,415 MB | 4.6x less than RustSASA |
| SwissProt | 157 MB | 1,131 MB | 2,187 MB | 7.2x less than RustSASA |
Memory advantage increases with dataset size. zsasa_bitmask has virtually no memory overhead compared to standard zsasa, while Lahuta Bitmask uses 5–14x more memory than zsasa_bitmask.
Bitmask Variants
The _bitmask variants use LUT (look-up table) bitmask neighbor lists instead of per-atom arrays:
- zsasa_bitmask: ~2.5–3x faster than regular zsasa, minimal memory overhead (~3 MB)
- Lahuta Bitmask: ~2.5–3x faster than regular Lahuta, but with significant memory overhead
- zsasa_bitmask is 1.3–1.6x faster than Lahuta Bitmask across all datasets
- Requires n_points ≥ 128
Datasets Larger Than Available RAM
zsasa uses mmap for file I/O. When the dataset fits in RAM, all file data stays in the OS page cache and performance depends purely on compute speed. When the dataset exceeds available RAM, mmap page faults must read from storage, and performance becomes storage I/O-bound regardless of compute efficiency.
In this regime, multiple worker threads fault pages in random order, so effective read throughput drops well below the SSD's sequential maximum. This is not specific to zsasa — any mmap-based tool (including Lahuta and RustSASA) hits the same bottleneck, and bitmask variants converge to similar wall-clock times (as seen in the M4 32 GB results).
zsasa's low memory footprint helps here: less RSS means more RAM is available for the OS page cache, which can reduce page fault frequency.
Methodology
Uses hyperfine for timing:
- Warmup runs (default 3) to eliminate cold-start effects
- Multiple timed runs for statistical reliability (3–10 runs)
- IQR outlier filtering applied when ≥5 runs
- Reports mean and stddev after filtering
Tool Configurations
| Tool | Configurations |
|---|---|
| zsasa (Zig) | f64/f32 precision, standard and bitmask variants |
| Lahuta (Zig) | Standard and bitmask variants |
| FreeSASA (C) | Sequential batch wrapper (sasa_batch.cpp) |
| RustSASA (Rust) | Native multi-threading |
Notes
- Test points: All benchmarks use 128 test points (required for Lahuta bitmask support).
- Input format: All tools use PDB input for fair comparison. zsasa supports JSON input which would be faster.
- FreeSASA: The CLI only supports single-threaded batch processing. A custom wrapper processes files sequentially.
- SwissProt (M4): I/O-bound due to mmap on 32 GB RAM — bitmask tools converge at ~11 min.
Running Benchmarks
# E. coli proteome (10 threads, 10 runs)
./benchmarks/scripts/bench_batch.py \
-i benchmarks/UP000000625_83333_ECOLI_v6/pdb \
-n ecoli --runs 10 --threads 10 -N 128
# Human proteome (10 threads, 10 runs)
./benchmarks/scripts/bench_batch.py \
-i benchmarks/UP000005640_9606_HUMAN_v6/pdb \
-n human --runs 10 --threads 10 -N 128
# SwissProt (large dataset, 3 runs)
./benchmarks/scripts/bench_batch.py \
-i /path/to/swissprot_pdb_v6 \
-n swissprot --runs 3 --threads 10 -N 128
Options
| Option | Description | Default |
|---|---|---|
--input, -i | Input directory (PDB files) | (required) |
--name, -n | Benchmark name | (required) |
--threads, -T | Thread counts: 1,8,10 or 1-10 | 1,8 |
--runs, -r | Number of benchmark runs | 3 |
--warmup, -w | Number of warmup runs | 3 |
--n-points, -N | Number of sphere test points | 100 |
--tool, -t | Tool: zig, zig_bitmask, freesasa, rustsasa, lahuta, lahuta_bitmask (repeatable) | all |
--output, -o | Output directory | results/batch/<N>/<name> |
--dry-run | Show commands without running | false |
Analysis
./benchmarks/scripts/analyze_batch.py summary -N 128 # Summary table
./benchmarks/scripts/analyze_batch.py summary -N 128 -n ecoli # Specific benchmark
./benchmarks/scripts/analyze_batch.py plot -N 128 -n ecoli # Time charts
./benchmarks/scripts/analyze_batch.py memory -N 128 -n ecoli # Memory charts
./benchmarks/scripts/analyze_batch.py all -N 128 # Everything
Related Documents
- Single-File Benchmarks — Per-structure performance across 2,013 structures
- MD Trajectory Benchmarks — Molecular dynamics trajectory analysis
- SASA Validation — Accuracy validation against FreeSASA