Single-File SASA Benchmarks
Large-scale benchmark results for zsasa using Shrake-Rupley algorithm.
- Dataset: 2,013 structures (stratified sampling from PDB + AlphaFold)
- Tools: zsasa (f64/f32, standard/bitmask), FreeSASA (C), RustSASA (Rust)
Note:
- Absolute execution times are environment-dependent. Relative speedup ratios are the meaningful metric for comparison.
- All implementations use identical parameters:
n_points=100,probe_radius=1.4Å- Benchmarks measure SASA calculation time only (file I/O excluded). FreeSASA and RustSASA have unstable I/O timing, so SASA-only measurement is used for fair comparison. See Methodology for details.
- SASA accuracy validated: mean error <0.001% vs FreeSASA reference. See validation for details.
TL;DR
zsasa's key advantage: Large structures + Multi-threading
| Speedup at threads=10 (50k+ atoms, n=794) | Thread Scaling (50k+ atoms) |
|---|---|
 |  |
Key Results (50k+ atoms, n=794, threads=10):
- 1.89x median speedup vs FreeSASA
- 1.84x median speedup vs RustSASA
- Bitmask mode: ~15x less memory with competitive speed on large structures
Test Environment
| Item | Value |
|---|---|
| Machine | MacBook Pro |
| Chip | Apple M4 |
| Cores | 10 (4 performance + 6 efficiency) |
| Memory | 32 GB |
| OS | macOS |
Overall Statistics (threads=10)
| Tool | Structures | Median (ms) | Mean (ms) | P95 (ms) |
|---|---|---|---|---|
| zsasa_f64 | 2,013 | 8.53 | 84.82 | 243.25 |
| zsasa_f32 | 2,013 | 8.75 | 83.68 | 243.75 |
| zsasa_f64_bitmask | 2,013 | 19.49 | 77.38 | 198.77 |
| zsasa_f32_bitmask | 2,013 | 19.28 | 76.34 | 196.30 |
| FreeSASA | 2,012 | 15.47 | 482.69 | 490.89 |
| RustSASA | 2,013 | 16.29 | 160.29 | 449.29 |
Note: zsasa_f64 has lower median but bitmask variants have lower mean/P95. This is because bitmask avoids the worst-case memory allocation overhead on very large structures, resulting in more stable performance at the high end.
FreeSASA shows 2,012 instead of 2,013 because it was OOM-killed (exit code 137) on the largest structure in the dataset —
9mjn(~3.8M atoms, 316MB PDB). All zsasa variants completed this structure successfully. See #247 for details.
Speedup by Structure Size (threads=10)
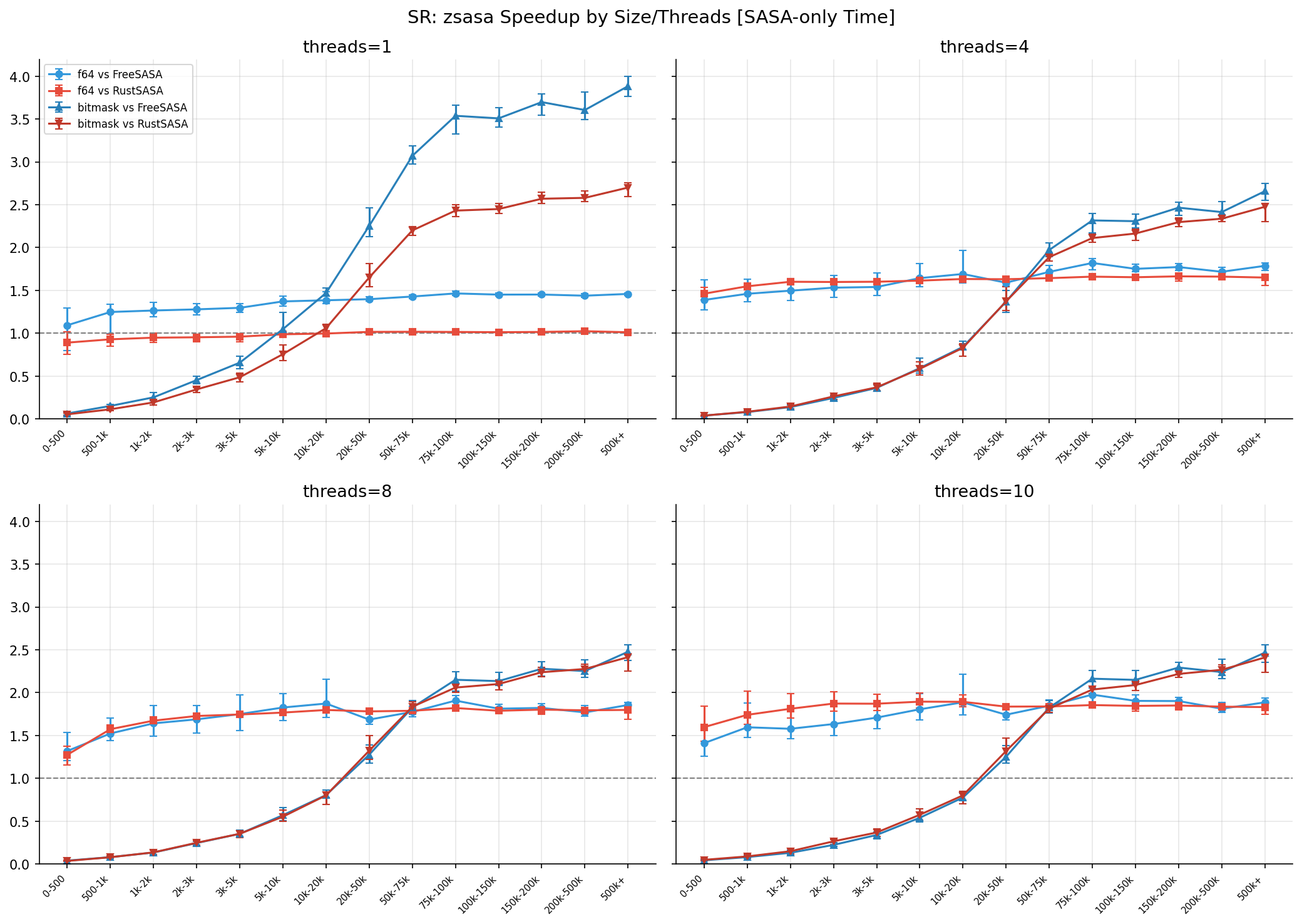
| Size Bin | Count | vs FreeSASA | vs RustSASA |
|---|---|---|---|
| 0-500 | 150 | 1.41x | 1.60x |
| 500-1k | 150 | 1.60x | 1.74x |
| 1k-2k | 150 | 1.58x | 1.81x |
| 2k-3k | 150 | 1.63x | 1.87x |
| 3k-5k | 150 | 1.71x | 1.87x |
| 5k-10k | 150 | 1.81x | 1.89x |
| 10k-20k | 150 | 1.89x | 1.89x |
| 20k-50k | 149 | 1.74x | 1.84x |
| 50k-75k | 150 | 1.85x | 1.84x |
| 75k-100k | 152 | 1.98x | 1.85x |
| 100k-150k | 148 | 1.90x | 1.85x |
| 150k-200k | 150 | 1.90x | 1.85x |
| 200k-500k | 150 | 1.83x | 1.84x |
| 500k+ | 63 | 1.89x | 1.83x |
Observations:
- vs FreeSASA: speedup increases with structure size, peaking at 1.98x (75k-100k)
- vs RustSASA: consistently 1.8-1.9x across medium-to-large structures
- Small structures (0-500): overhead dominant, but still 1.4-1.6x faster
Single-Thread Performance (threads=1)
Single-threaded comparison (excluding parallelization effects):
| Size Bin | Count | vs FreeSASA | vs RustSASA |
|---|---|---|---|
| 0-500 | 150 | 1.09x | 0.89x |
| 500-1k | 150 | 1.25x | 0.93x |
| 1k-2k | 150 | 1.27x | 0.95x |
| 2k-3k | 150 | 1.28x | 0.95x |
| 3k-5k | 150 | 1.30x | 0.96x |
| 5k-10k | 150 | 1.37x | 0.99x |
| 10k-20k | 150 | 1.38x | 1.00x |
| 20k-50k | 149 | 1.40x | 1.02x |
| 50k-75k | 150 | 1.43x | 1.02x |
| 75k-100k | 152 | 1.46x | 1.01x |
| 100k-150k | 148 | 1.45x | 1.01x |
| 150k-200k | 150 | 1.45x | 1.02x |
| 200k-500k | 150 | 1.44x | 1.02x |
| 500k+ | 63 | 1.46x | 1.01x |
Observations:
- vs FreeSASA: 1.4-1.5x on large structures (SIMD optimization)
- vs RustSASA: nearly equal at t=1 (both use SIMD), zsasa slightly ahead on large structures
- The gap between t=1 and t=10 speedups shows zsasa's parallel efficiency advantage
Thread Scaling
Median Execution Time by Thread Count

| Threads | zsasa_f64 (ms) | FreeSASA (ms) | RustSASA (ms) | zsasa_f64_bitmask (ms) |
|---|---|---|---|---|
| 1 | 23.11 | 30.84 | 22.63 | 20.98 |
| 4 | 10.32 | 16.73 | 16.82 | 19.80 |
| 8 | 9.00 | 15.95 | 16.07 | 19.60 |
| 10 | 8.53 | 15.47 | 16.29 | 19.49 |
Speedup from threads=1 to threads=10:
- zsasa_f64: 23.11 → 8.53 = 2.71x
- FreeSASA: 30.84 → 15.47 = 1.99x
- RustSASA: 22.63 → 16.29 = 1.39x
Key Insight:
- zsasa_f64 scales best with thread count, maintaining large gains up to t=10
- RustSASA barely improves beyond t=1 (parallel efficiency issue)
- Bitmask variants have low per-structure overhead but limited thread scaling (work is already cheap per atom)
Bitmask Variants
Bitmask mode uses LUT bitmask neighbor lists instead of full float arrays. This dramatically reduces memory usage at the cost of slightly higher per-structure computation time and minor accuracy differences.
When to Use
| Mode | Best For | Trade-off |
|---|---|---|
| Standard (f64) | Maximum speed, highest accuracy | Higher memory usage |
| Bitmask (f64) | Large structures, memory-constrained | ~15x less memory for 500k+ atoms; minor accuracy difference |
Accuracy note: Bitmask mode uses a fixed cutoff for neighbor detection, which may produce slightly different SASA values compared to standard mode. The difference is negligible for most use cases. See SASA Validation for detailed accuracy analysis.
Overall Performance (threads=10)
| Tool | Median (ms) | Mean (ms) | P95 (ms) |
|---|---|---|---|
| zsasa_f64 | 8.53 | 84.82 | 243.25 |
| zsasa_f64_bitmask | 19.49 | 77.38 | 198.77 |
| zsasa_f32 | 8.75 | 83.68 | 243.75 |
| zsasa_f32_bitmask | 19.28 | 76.34 | 196.30 |
Large Structure Performance (100k+ atoms, threads=10)
On large structures, bitmask variants close the speed gap and become competitive:
| Tool | n | Median (ms) | Mean (ms) | P95 (ms) |
|---|---|---|---|---|
| zsasa_f64 | 512 | 147.25 | 285.98 | 1,051.50 |
| zsasa_f64_bitmask | 512 | 123.95 | 230.15 | 817.08 |
| zsasa_f32 | 512 | 145.25 | 282.09 | 1,038.91 |
| zsasa_f32_bitmask | 512 | 121.90 | 227.25 | 806.76 |
Key finding: For 100k+ atom structures, bitmask is actually faster than standard mode (16% lower median, 20% lower mean) while using far less memory. The bitmask's fixed-size neighbor storage avoids the allocation overhead that dominates large structures.
Memory Comparison
Peak memory (RSS) measured by hyperfine (threads=1):
| Memory vs Structure Size | Memory Scatter |
|---|---|
 |  |
Observations:
- FreeSASA uses the most memory across all sizes (~15x more than zsasa for 500k+ atoms)
- RustSASA uses ~5x more than zsasa for large structures
- zsasa standard and bitmask have similar memory for small structures
- For 500k+ atoms, bitmask uses significantly less memory than standard mode
Large Structure Analysis
Summary (50k+ atoms, n=794, threads=10)
| Speedup at threads=10 | Thread Scaling |
|---|---|
 |  |
| Comparison | Median Speedup |
|---|---|
| vs FreeSASA | 1.89x |
| vs RustSASA | 1.84x |
Best Speedup Structures (50k+ atoms)
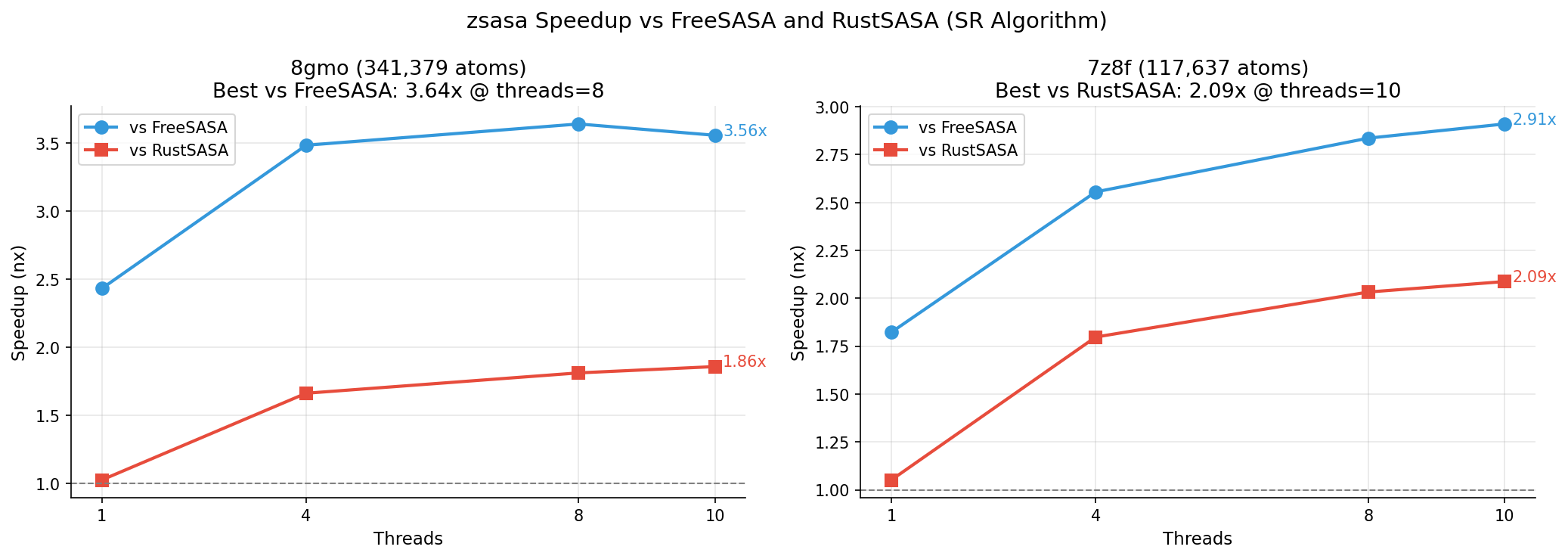
Maximum Structure Thread Scaling

Execution Time Distribution
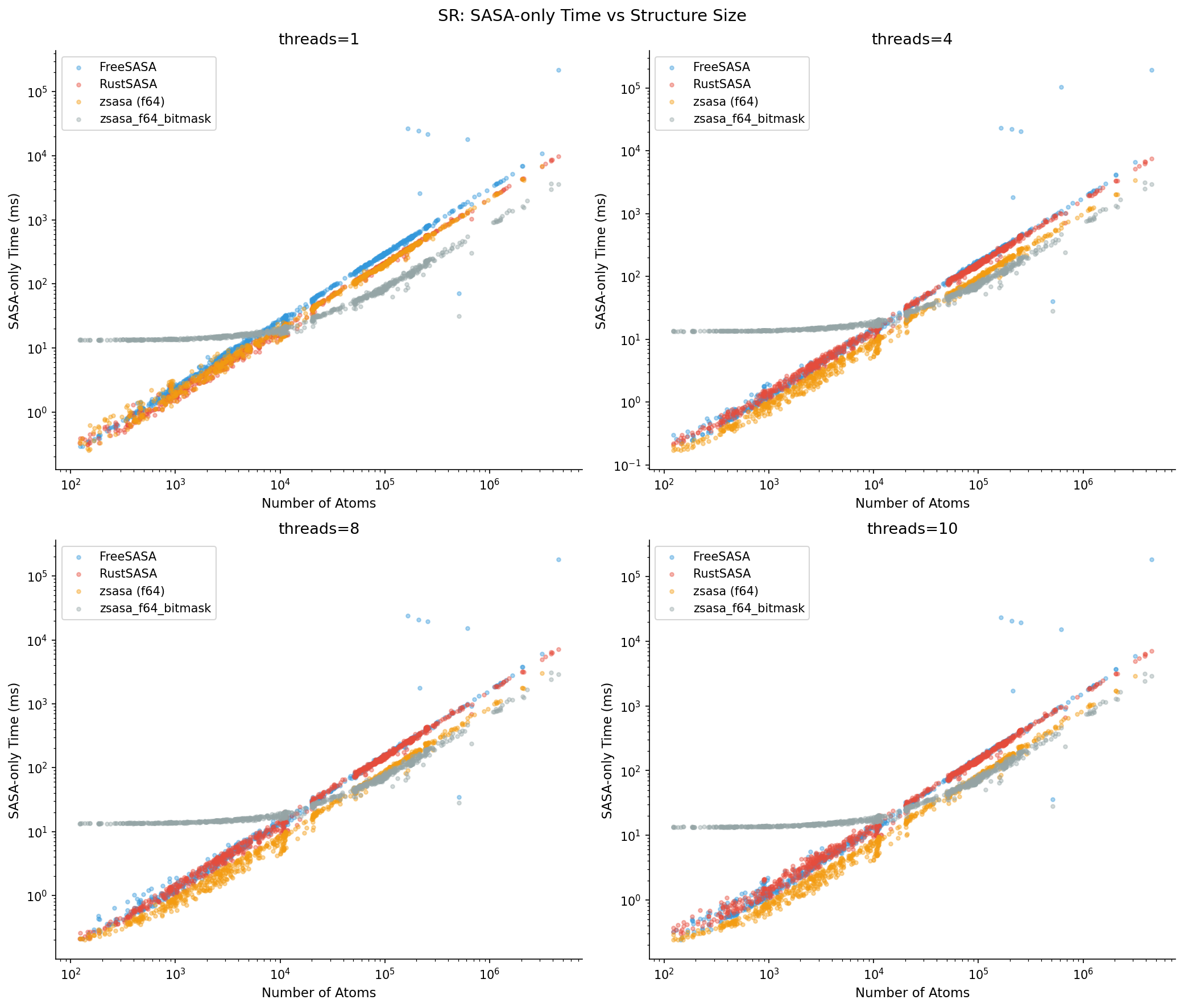
Observations:
- Nearly linear on log scale → O(N) neighbor list is effective (all tools use cell list)
- zsasa (orange) is consistently lower (faster) across all sizes
- Gap between tools widens with increasing thread count
- Few outliers → stable performance
Stability: Outlier Structures
One of zsasa's key advantages is consistent, predictable performance across all structures. FreeSASA and RustSASA exhibit pathological slowdowns on certain structures, where computation time spikes by orders of magnitude. zsasa shows no such behavior.
FreeSASA Pathological Cases

19 structures where FreeSASA takes 3x–114x longer than zsasa (SASA-only time, threads=1). The worst case (7n9f, 212,962 atoms) shows 113.9x slowdown. The cause is unknown but appears structure-dependent.
RustSASA Pathological Cases
Example: 9gdy (509,160 atoms) — RustSASA takes ~14,000ms vs zsasa's ~50ms at threads=1:

RustSASA is ~280x slower on this structure, and the slowdown persists across all thread counts. Including wall-clock (I/O) timing reveals even more outlier structures for both FreeSASA and RustSASA.
Key takeaway: zsasa produces stable, predictable timing across all 2,013 structures tested. No pathological cases were observed. This predictability is important for batch processing and pipeline reliability.
FreeSASA OOM on Very Large Structures
FreeSASA was OOM-killed (SIGKILL, exit code 137) when processing 9mjn (~3.8M atoms), the largest structure in the dataset. This was the only failure across all 2,013 structures. All zsasa variants (f64, f32, standard, bitmask) completed this structure without issue, demonstrating zsasa's ability to handle arbitrarily large inputs within bounded memory. (#247)
Per-Bin Sample Results
Thread scaling details on representative structures selected from each size bin.
| Bin | Atom Range | Sample Plot |
|---|---|---|
| 0-500 | 0 – 500 | View |
| 500-1k | 500 – 1,000 | View |
| 1k-2k | 1,000 – 2,000 | View |
| 2k-3k | 2,000 – 3,000 | View |
| 3k-5k | 3,000 – 5,000 | View |
| 5k-10k | 5,000 – 10,000 | View |
| 10k-20k | 10,000 – 20,000 | View |
| 20k-50k | 20,000 – 50,000 | View |
| 50k-75k | 50,000 – 75,000 | View |
| 75k-100k | 75,000 – 100,000 | View |
| 100k-150k | 100,000 – 150,000 | View |
| 150k-200k | 150,000 – 200,000 | View |
| 200k-500k | 200,000 – 500,000 | View |
| 500k+ | 500,000+ | View |
Key Takeaways
Why is zsasa faster? SIMD optimization (8-wide distance calculation), multi-threading with work stealing, and spatial hashing for O(1) neighbor lookup.
-
Consistent advantage across all sizes
- 1.9x vs FreeSASA, 1.8x vs RustSASA on large structures (threads=10)
- Outperforms FreeSASA at all sizes (even 0-500 atoms)
-
Best thread scaling
- 2.71x scaling from t=1 to t=10 (vs 1.99x FreeSASA, 1.39x RustSASA)
-
Memory-efficient bitmask mode
- ~15x less memory for 500k+ atom structures
- Competitive speed with lower P95 latency
-
Accurate results
- See validation for detailed accuracy analysis
Methodology
SASA-Only Timing
For fair comparison, we measure SASA calculation time only. File I/O is excluded because FreeSASA and RustSASA exhibit unstable I/O timing.
Total time = File I/O + SASA calculation + Output
^^^^^^^^^^^^^^^^
Only this is measured
Measurement method for each implementation:
| Implementation | Method |
|---|---|
| zsasa (all variants) | Internal measurement via --timing option (stderr output) |
| FreeSASA C | Patched binary outputs SASA calculation time to stderr |
| RustSASA | Patched binary outputs SASA_TIME_US to stderr |
Parameters
| Parameter | Value | Notes |
|---|---|---|
| Algorithm | Shrake-Rupley | Supported by all implementations |
| n_points | 100 | Number of test points |
| probe_radius | 1.4 Å | Water molecule radius |
| Warmup | 1 | Discarded before measurement |
| Runs | 3 | Average value used |
Tool Variants
| Tool | Precision | Neighbor Storage | Notes |
|---|---|---|---|
| zsasa_f64 | f64 | Full array | Default, highest accuracy |
| zsasa_f32 | f32 | Full array | ~3% faster than f64 at t=1 |
| zsasa_f64_bitmask | f64 | Bitmask | ~15x less memory for 500k+ atoms |
| zsasa_f32_bitmask | f32 | Bitmask | Lowest memory usage |
| FreeSASA | f64 | — | C reference implementation |
| RustSASA | f32 | — | Rust implementation |
Stratified Sampling
Stratified sampling from PDB and AlphaFold structures, ~150 structures per bin (14 bins):
| Bin | Atom Range | Count |
|---|---|---|
| 0-500 | 0 – 500 | 150 |
| 500-1k | 500 – 1,000 | 150 |
| 1k-2k | 1,000 – 2,000 | 150 |
| 2k-3k | 2,000 – 3,000 | 150 |
| 3k-5k | 3,000 – 5,000 | 150 |
| 5k-10k | 5,000 – 10,000 | 150 |
| 10k-20k | 10,000 – 20,000 | 150 |
| 20k-50k | 20,000 – 50,000 | 149 |
| 50k-75k | 50,000 – 75,000 | 150 |
| 75k-100k | 75,000 – 100,000 | 152 |
| 100k-150k | 100,000 – 150,000 | 148 |
| 150k-200k | 150,000 – 200,000 | 150 |
| 200k-500k | 200,000 – 500,000 | 150 |
| 500k+ | 500,000+ | 63 |
| Total | 2,013 |
Running Benchmarks
Setup
# Build Zig binary
zig build -Doptimize=ReleaseFast
# External tools setup (for comparison)
cd benchmarks/external
git clone https://github.com/N283T/freesasa-bench.git
cd freesasa-bench && ./configure --enable-threads && make && cd ..
git clone --recursive https://github.com/N283T/rustsasa-bench.git
cd rustsasa-bench && cargo build --release --features cli && cd ..
Index & Sample Generation
# Create index (first time only)
./benchmarks/scripts/build_index.py benchmarks/inputs
# Check distribution
./benchmarks/scripts/sample.py benchmarks/inputs/index.json --analyze
# Generate sample
./benchmarks/scripts/sample.py benchmarks/inputs/index.json \
--seed 42 \
-o benchmarks/dataset/sample.json
Running
# Basic usage
./benchmarks/scripts/bench.py --tool zig --algorithm sr --threads 1,4,8,10
# All tools
./benchmarks/scripts/bench.py --tool zig --algorithm sr --threads 1,4,8,10
./benchmarks/scripts/bench.py --tool freesasa --algorithm sr --threads 1,4,8,10
./benchmarks/scripts/bench.py --tool rustsasa --algorithm sr --threads 1,4,8,10
# With bitmask mode (Zig only)
./benchmarks/scripts/bench.py --tool zig --algorithm sr --bitmask --threads 1,4,8,10
# With f32 precision (Zig only)
./benchmarks/scripts/bench.py --tool zig --algorithm sr --precision f32 --threads 1,4,8,10
Analysis & Visualization
# Summary tables (SASA-only metric, default)
./benchmarks/scripts/analyze.py summary
# Summary tables (wall-clock metric)
./benchmarks/scripts/analyze.py summary --metric wall
# Generate all plots
./benchmarks/scripts/analyze.py all
# Individual plot types
./benchmarks/scripts/analyze.py scatter # Atoms vs time scatter
./benchmarks/scripts/analyze.py threads # Thread scaling
./benchmarks/scripts/analyze.py grid # Speedup grid by size/threads
./benchmarks/scripts/analyze.py samples # Per-bin sample plots
./benchmarks/scripts/analyze.py large # Large structure analysis
./benchmarks/scripts/analyze.py memory # Peak memory comparison
./benchmarks/scripts/analyze.py speedup # Best speedup structures
./benchmarks/scripts/analyze.py outliers # Outlier advantage plots
# Export to CSV
./benchmarks/scripts/analyze.py export-csv
Scripts
| Script | Purpose |
|---|---|
build_index.py | Create atom count index from all input files |
sample.py | Stratified sampling from index |
bench.py | Run benchmarks (single-file mode) |
analyze.py | Analyze results and generate plots |
Notes
- Initial runs are slow: due to file cache and warmup effects
- Thread count depends on CPU: optimal when matching physical core count
- External tools require patches: SASA-only timing requires modified binaries
Appendix: Lee-Richards (LR) Algorithm
zsasa also supports the Lee-Richards (LR) algorithm, which uses slice-based numerical integration instead of sphere test points.
Key Differences from Shrake-Rupley
| Property | Shrake-Rupley (SR) | Lee-Richards (LR) |
|---|---|---|
| Method | Test points on sphere | Slice integration |
| Speed | Faster | 3-5x slower |
| Accuracy | Depends on n_points | Depends on n_slices |
| Supported by | zsasa, FreeSASA, RustSASA | zsasa, FreeSASA |
LR benchmarks are not included in this comparison because RustSASA does not support LR, making a three-way comparison impossible. For LR-specific analysis, see the analyze_lr.py script.
# Run LR benchmark
./benchmarks/scripts/bench_lr.py --tool zig --algorithm lr --n-slices 20
# Analyze LR results
./benchmarks/scripts/analyze_lr.py summary
Related Documents
- Batch Processing Benchmarks — proteome-scale batch processing
- MD Trajectory Benchmarks — molecular dynamics trajectory analysis
- SASA Validation — accuracy validation vs FreeSASA reference
- Algorithms — algorithm comparison and usage